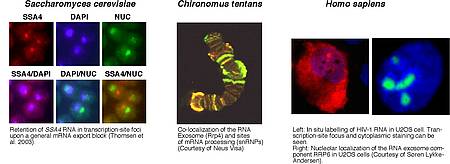
Eukaryotic RNAs are produced and matured in the cell nucleus. Major events in the biogenesis of most RNAs, including splicing, 3’ end processing and RNP formation, occurs co-transcriptionally. Coupling with transcription increases the efficiency and specificity of the processing-packaging systems and sets up an assembly line with “built in” quality control (QC). This is because enzymes and complexes harboring RNA degradative activity are immediately available to rapidly degrade the RNAs of aberrant, or sub-optimal RNP molecules. The objective of this proposal is to elucidate the mechanisms governing nuclear RNA surveillance. Transcription, processing and degradation of RNA have traditionally been studied as independent processes, and each of the participating groups is an expert in one (or a few) of these. To break new scientific ground pooling of expertise, tools and methodologies from different fields and model organisms is needed.
PROJECT LEADER:
Dr. Torben Heick Jensen
University of Aarhus
Department of Molecular Biology
8000 Aarhus, Denmark
Phone: +45 8942 2609
Fax: +45 8619 6500
website: http://dna8.mb.au.dk/group/thj/
PRINCIPAL INVESTIGATORS:
Andrès Aguilera, University of Sevilla, ES
Irene Bozzoni, CNR-IBPM, University of Rome “La Sapienza”, IT
Alain Jacquier, Institut Pasteur, CNRS, FR
Domenico Libri, CNRS-CGM, FR
Nicholas Proudfoot, Oxford University, UK
David Tollervey, University of Edinburgh, UK
Neus Visa, Stockholm University, SE